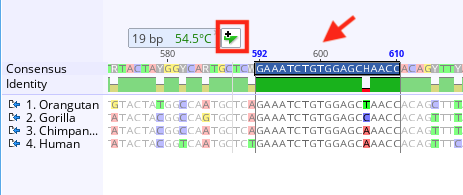
Note that for performance reasons these statistics are only calculated on documents comprised of less than 10,000 bp or aa (this threshold applies to the total number of residues and gaps over all sequences in a document). If you wish to add these statistics to the document table for documents created in earlier versions of Geneious, select the folder containing the document and go to Tools → Preferences → Appearance and Behavior and select Recalculate statistics now. Molecular weight and protein statistics were added to the document table in Prime 2021, and will not appear in the table for documents created in an earlier version of Geneious unless that document is edited in Prime 2021. The value in the document table will be for the entire document, not the currently selected region. These include sequence length, % pairwise identity, % identical sites, mean coverage, molecular weight and several protein statistics such as extinction coefficient and isoelectric point. Several of the metrics displayed in the Statistics tab can also be displayed as columns in the document table. The length and number of sequences currently selected is shown at the top of the Statistics tab. 
If only part of the sequence/alignment or assembly is selected then the statistics displayed will correspond to the highlighted part. “agents” and collaboration modes) - the best advice I can give is to download it, do the tutorial and test drive it on a bioinformatics task that you regularly conduct - it won't disappoint.The Statistics tab displays statistics about the sequence(s) being viewed. 
The software almost has too many features to comment on in any one review (e.g. Based on the hours of time this program has saved me already I would not even blink at the academic price tag of $133 (USD) - or cheaper for students. The Geneious team has made available a freeware version and a pro version that (for a price) allows you to access more functionality. Moreover, the software is structured in such a way that “new” plugins can be written and incorporated with relative ease. My request to provide a variety of nucleotide colour schemes was implemented within 2 weeks. As is the case with any new software there are a few features that don't work quite as well as you might like - but the good news is that if you write to them they will likely fix it up asap. With the integration of sequence chromatograph viewing and editing of DNA sequences/alignments (new in version 2) Geneious is now firmly embedded in my research practices. If you are looking for a piece of bioinformatics software for teaching purposes look no further than Geneious. Students did a Genbank search, downloaded files, did an alignment and built a phylogenetic tree (with bootstrapping) all in the blink of an eye. Earlier this year I used the Beta-release in some undergraduate teaching labs - with great success. While the software has some “bell and whistles” for more complex tasks one of its key attributes is a simple user interface. The user interface of Geneious is both attractive and intuitive. The software logo “research in a flash” accurately describe the potential of Geneious - only months after its release it has become the “Swiss army knife” of bioinformatics software. Current and future releases of Geneious (offer a solution to (most) of these frustrations. Chopping and changing between different file formats and at times different operating systems (Mac, PC, Unix/Linux) has been a frustrating and time-consuming exercise. For a long time scientists who generate or analyse DNA sequences have been using multiple pieces of software to: 1) view chromatographs, 2) trim sequences 3) Blast data 4) align sequences 5) build phylogenetic trees. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
